 |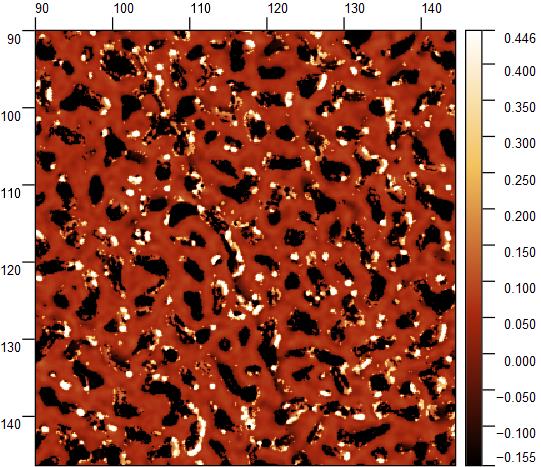  | 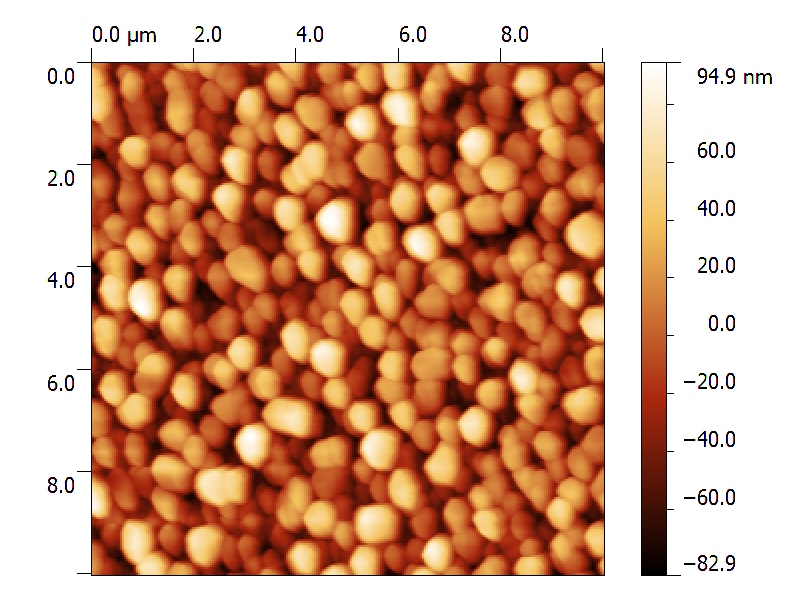 | 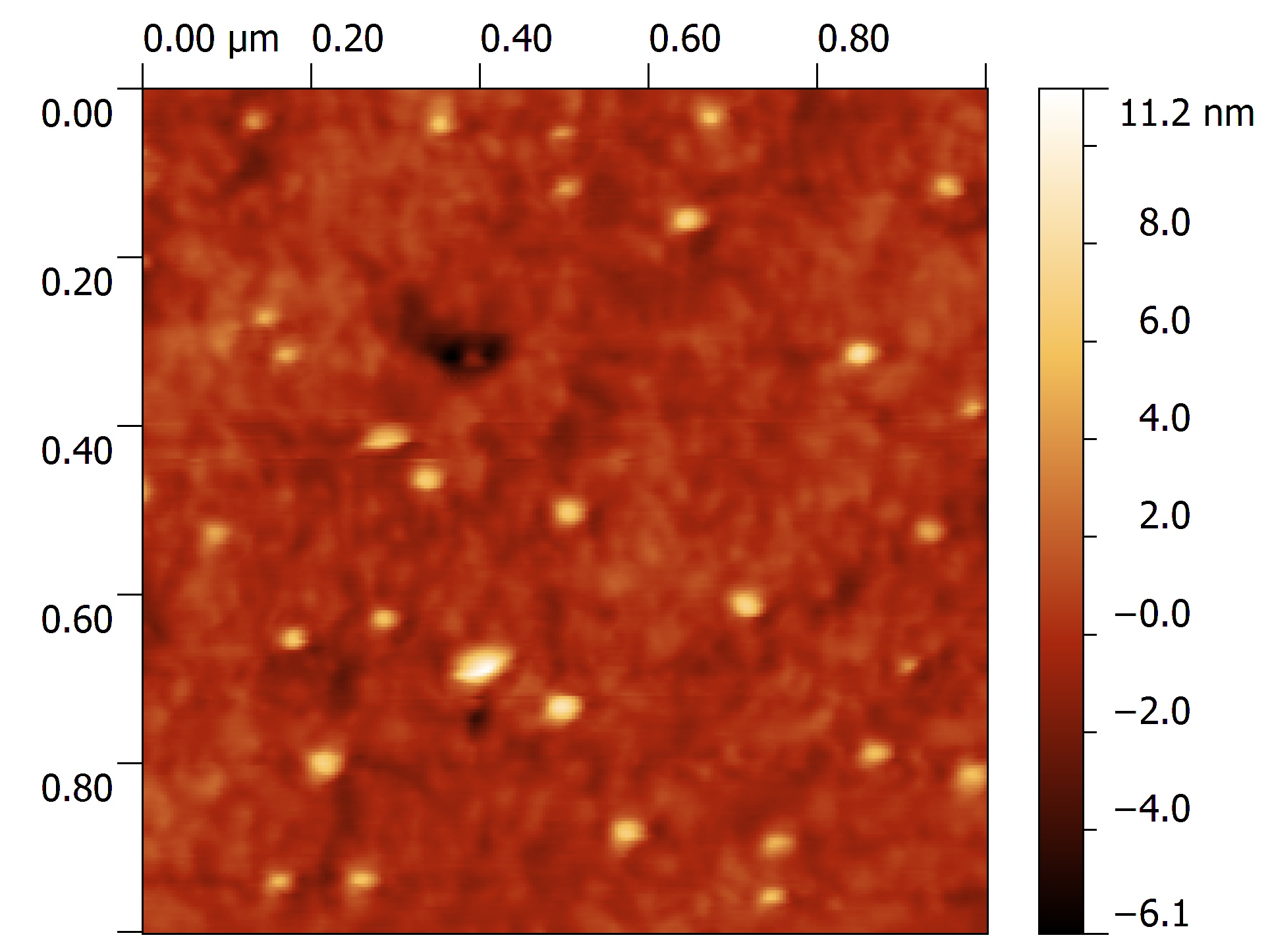 |
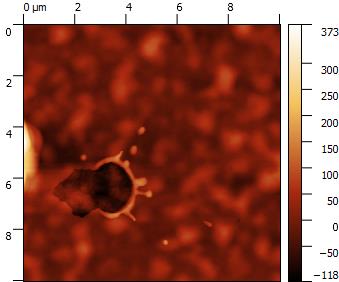
AFM data are (most of the times) surface or profile data.
%load_ext autoreload
%autoreload 2
import os
import numpy as np
from matplotlib import pyplot as plt
from traitlets.config.manager import BaseJSONConfigManager
from pathlib import Path
path = Path.home() / ".jupyter" / "nbconfig"
cm = BaseJSONConfigManager(config_dir=str(path))
cm.update(
"rise",
{
"theme": "sky",
"transition": "slide",
"start_slideshow_at": "selected",
"width": "90%",
"display": "inline-block",
"font-size": "0.6em",
"line-height": "1.2em",
"vertical-align": "top",
"header": "<h3>HNSFE 2020 - 2020/09/25</h3>",
"footer": "<h3>V. Cotroneo - INAF/Osservatorio Astronomico di Brera</h3>"
}
)
{'theme': 'sky',
'transition': 'slide',
'start_slideshow_at': 'selected',
'display': 'inline-block',
'vertical-align': 'top',
'width': '90%',
'font-size': '0.6em',
'line-height': '1.2em',
'header': '<h3>HNSFE 2020 - 2020/09/25</h3>',
'footer': '<h3>V. Cotroneo - INAF/Osservatorio Astronomico di Brera</h3>'}
 |  |  |  |
AFM data are (most of the times) surface or profile data.
Surface and Profile data (I will consider them as a subset) are of general interest in almost any field of science and technology, from optics, to microscopy, to material science and biology.
Many operations are common to different disciplines and instruments.
Data represent a map over a 2D grid (or more in general a set of points in the 3D space), where elevation, intensity, or other quantities are expressed as a function of 2D position, identified on surface by its relation with coordinate axis.
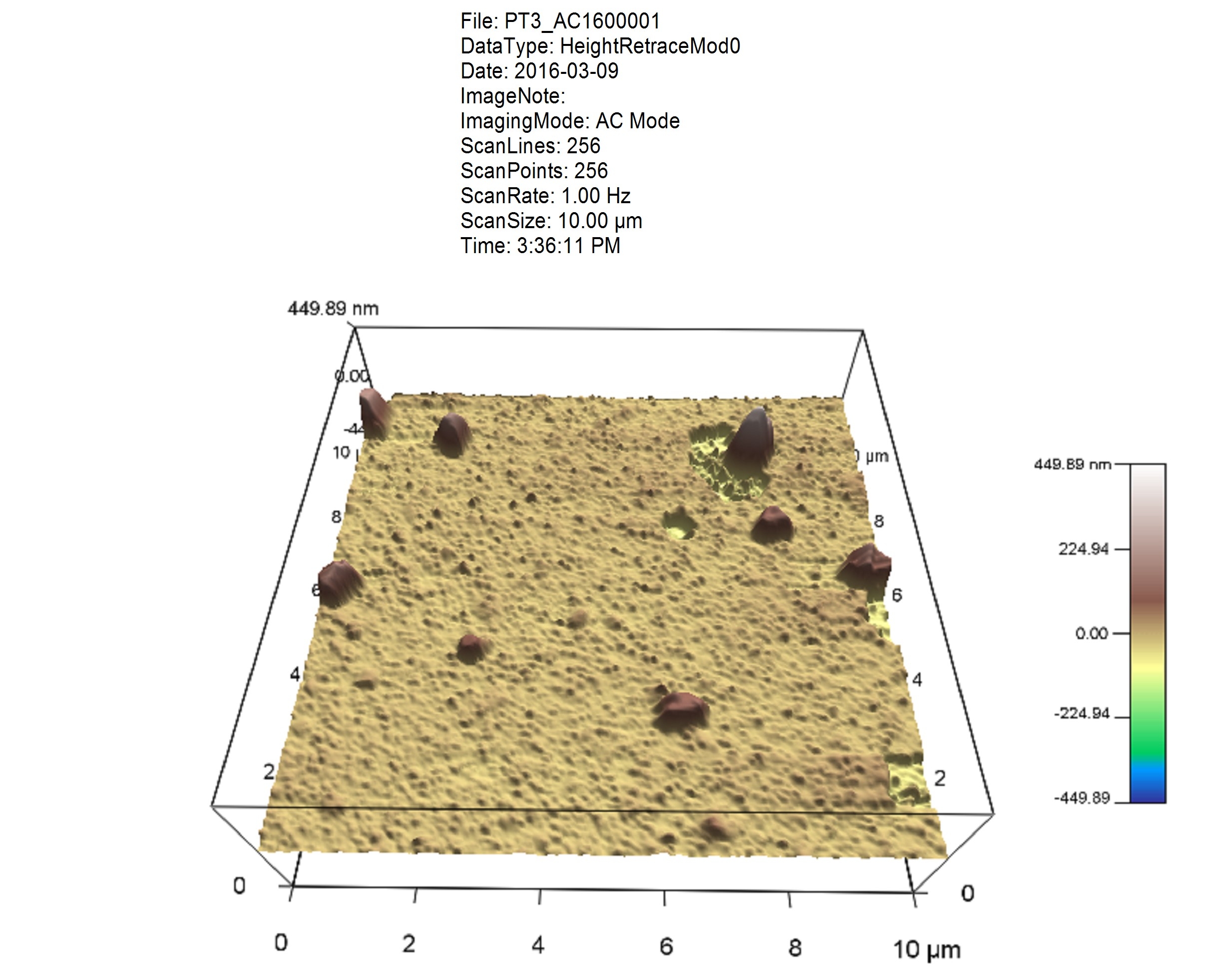

A set of operations (many of which are common in different disciplines, e.g. leveling, feature recognition, profile extraction) are performed on data.
Furthermore:
{'theme': 'sky', 'transition': 'slide', 'start_slideshow_at': 'selected', 'width': '90%', 'height': '100%', 'display': 'inline-block', 'font-size': '0.6em', 'line-height': '1.2em', 'vertical-align': 'top', 'header': '
Usually provided with the instrument. Typically GUI-based:
(*) Many of these tools implement some form of scripting (often Python based) or macro procedures, internally (scripting engine embedded in software) or externally (access to internal functions given through APIs od DLLs).
 |
 |
|
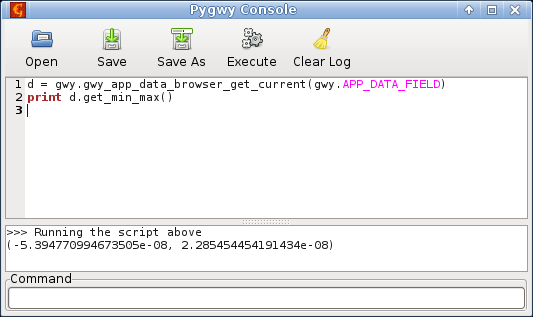
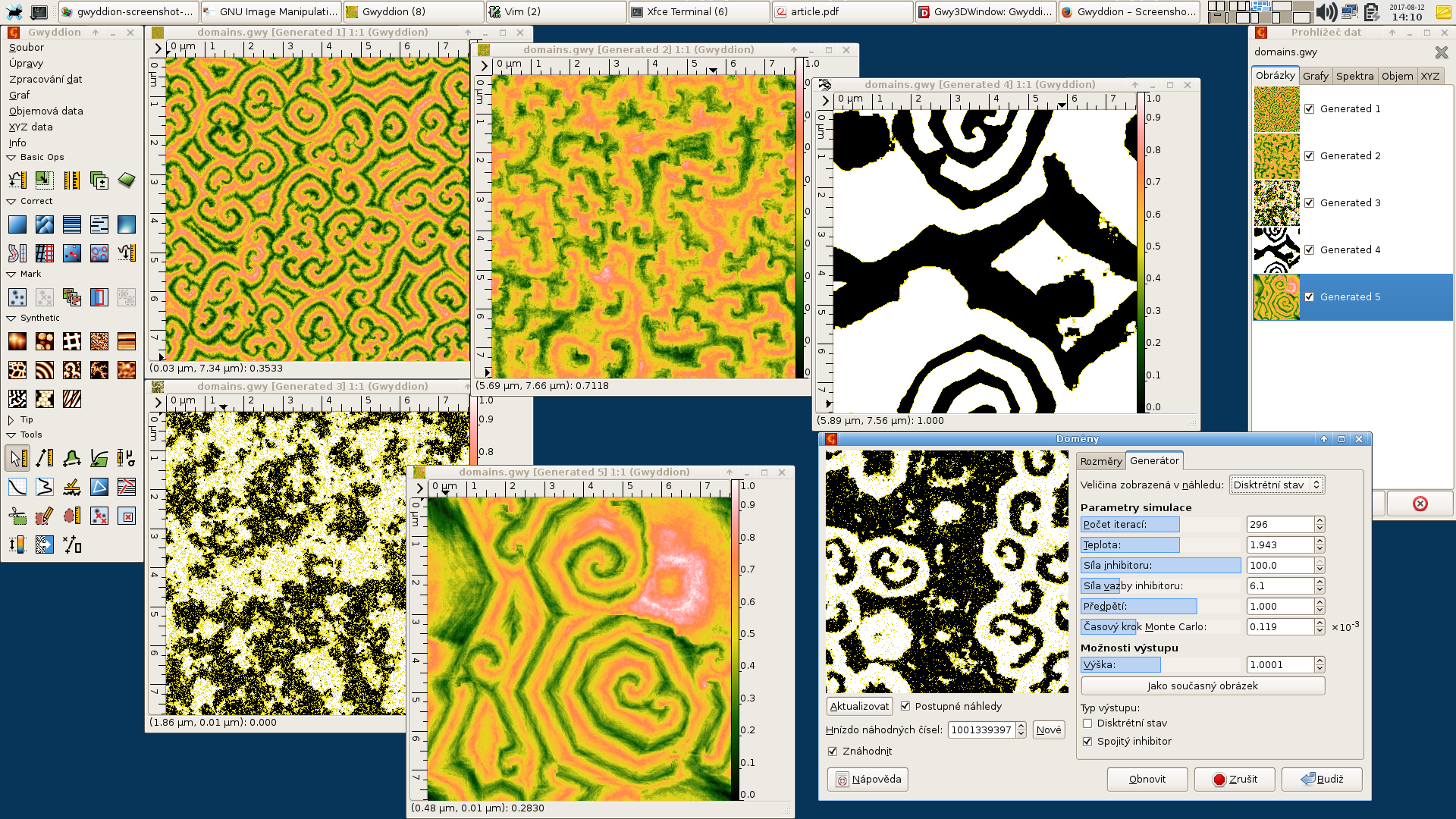
Gwyddion, developed for nanometrology, and probably the most successful software of this kind, is versatile and can handle many formats of surface data. It offers open source and has some form of interfacing with python, but is still based on a very proprietary graphical interface. |
 |
 |
Everybody writes their own software:
|
In many fields of science, the Python programming language, with its approach aimed at reusability, inverted this tendency: in the last years, several Python packages, tools and libraries emerged and established themselves as standard tools, bringing the language to be one of the most popular in science, if not the most popular.
 | 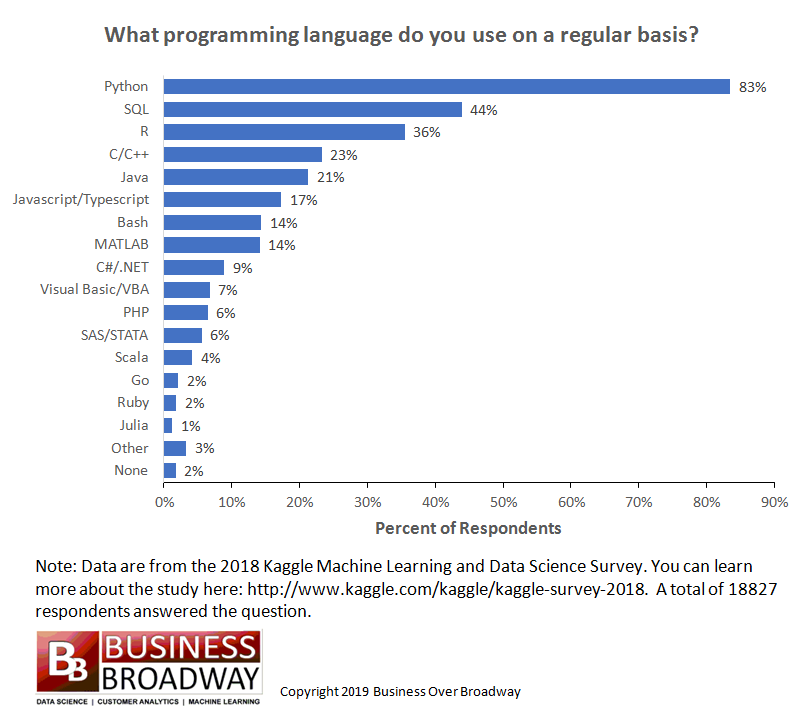 |
The result is an enormous base of users.
High quality libraries for practically any field of science.
Python is likely also the most commonly used language for scripting of surface-metrology instruments proprietary software.

However, to the knowledge of the author, there is not an universally recognized framework in any language and especially not in Python.
Python has good facility to deal with multidimensional arrays, but it lack the coordination with axis (It basically treat everything as a matrix).
The library was created during research on X-ray mirrors (X-ray mirrors are characterized at all wavelength from AFM to LTP, meaning from 1 um to meters scan size, profile or surface).
The pyXsurf library consists in a set of core routines and classes, with a quite uniform and well defined interface, which can internally handle coordinate transformations, resulting in much more intuitive operations and enabling complex actions on data compared to pure Python.
# {"backimage": "material\logos\logos.png"}
{'backimage': 'material\\logos\\logos.png'}
Python offers:
The implementation of this project in Python, gives to the library the following advantages:
Use cases and applicability of the program:
A generic example of use, applicable to several fields of application, can be the alignment and subtraction of two images, as shown in fig. 2 as handled by pySurf, for a case in which the alignment is interactively determined on the base of a separate set of data. The case is not trivial to handle with common software: the user needs to enter and exit several menus in the software GUI, save fiducials, retrieve them, and open and close several files. This is a set of actions that, when possible, is performed differently and with different limitations, in each software. The use of different softwares, also results in a non uniform output.
The library is mostly made of classes, of which the most important is Data2D, representing a measured surface. A Data2D object p1 can be created by passing data, e.g.:
In order to illustrate pyXsurf approach and its relevance to different contexts in the field of surface data analysis, it can be useful to provide some minimal examples of its usage and syntax. The library is mostly made of classes, of which the most important is Data2D, representing a measured surface.
A Data2D object p1 can be created by passing data, e.g.:
p1 = Data2D(data,x,y)# This is just used to define data, x, y from data file
from pySurf.readers.instrumentReader import matrix_reader
f = r'material\AFM_images\M12 no corona 50 nc.dat'
data,x,y = matrix_reader(f,delimiter='',skip_header=4)
from pySurf.data2D_class import Data2D
D = Data2D(data,x,y,units=['um','um','nm'],scale=[1,1,1.e9],name=os.path.basename(f)) # data,x,y previously read from .dat file
We can do a basic plot.
D.plot(stats=1)
<Axes: title={'center': 'M12 no corona 50 nc.dat'}, xlabel='X (um)', ylabel='Y (um)'>
The units are already set up. Several options are available for customization, plot method follows standard plt.plot interface, the returned plot is an plt.Axes object, modifiable with usual commands.
Leveling can be performed with a number of options to level to same or different orders in x and y, or by line along each axis:
D.level?
Signature: D.level(*args, **kwargs) Docstring: level_data(data, x=None, y=None, degree=1, axis=None, byline=False, fit=False, *args, **kwargs) Use RA routines to remove degree 2D legendres or levellegendre if leveling by line. Degree can be scalar (it is duplicated) or 2-dim vector. must be scalar if leveling by line. Note the important difference between e.g. ``degree = 2`` and ``degree = (2,2)``. The first one uses degree as total degree, it expands then to xl,yl = [0,1,0,1,2,0],[0,0,1,1,0,2]. The second leveling by line (controlled by axis keyword) also handle nans. x and y are not used, but maintained for interface consistency. fit=True returns fit component instead of residuals File: c:\users\kovor\documents\python\pyxtel\source\pysurf\data2d_class.py Type: method
Leveling operations return a new object, Commmands can be chained, here plot method is called on the object resulting from D.level() (default remove plane):
plt.figure(2)
D.level().plot(stats=1) # remove plan, stats controls the info box
<Axes: title={'center': 'M12 no corona 50 nc.dat'}, xlabel='X (um)', ylabel='Y (um)'>
Statistical functions (same interface as plt.hist()):
D.level().histostats(); # data after plane removal
The distribution is quite irregular, we can look for a more advanced leveling.
Results can be assigned to variables, here Dl is the result of removing a plan and then second order Legendre in horizontal direction for each line is removed:
plt.figure(3)
Dl = D.level().level(2,axis=1)
Dl.plot(stats=1)
<Axes: title={'center': 'M12 no corona 50 nc.dat'}, xlabel='X (um)', ylabel='Y (um)'>
Several common operations, remove outliers:
plt.figure()
Dl.remove_outliers(nsigma =3).plot()
<Axes: title={'center': 'M12 no corona 50 nc.dat'}, xlabel='X (um)', ylabel='Y (um)'>
Dl.remove_outliers(nsigma=3).histostats();
Dl.plot()
<Axes: title={'center': 'M12 no corona 50 nc.dat'}, xlabel='X (um)', ylabel='Y (um)'>
We can try to remove outliers and level again by line (brings very close to data intrinsic errors):
Dl = D.level().level(2,axis=1).remove_outliers(nsigma =3).level(2,axis=1)
Dl.plot()
<Axes: title={'center': 'M12 no corona 50 nc.dat'}, xlabel='X (um)', ylabel='Y (um)'>
Dl.histostats();
Method Data2D.extract_profile profiles can be extracted as Profile objects according to different criteria, including interactive mode, by point and click. In this case horizontal profiles are extracted at fixed ys.
from pyProfile.profile_class import Profile
ypos = [50,100,150,200]
plist = [] # make a list of profiles
for y in ypos: # populate the list for the positions in y using `Data2D.extract_profile`.
plist.append(Profile(*Dl.extract_profile([10,y],[240,y],along=True),name = y))
for pp in plist:
pp.level(zero='top').plot()
plt.grid()
plt.legend(loc=0)
<matplotlib.legend.Legend at 0x28c47d895b0>
ps = Dl.psd(analysis=True)
WARNING: low limit detected in prange, can cause problems with log color scale.
<Figure size 640x480 with 0 Axes>
ps.plot()
<Axes: title={'center': '2D PSD'}, xlabel='X (um)', ylabel='freq. (um$^{-1}$)'>
from pyProfile.psd import plot_psd
plot_psd(*ps.avgpsd())
No handles with labels found to put in legend.
Applicable cases in my experience:

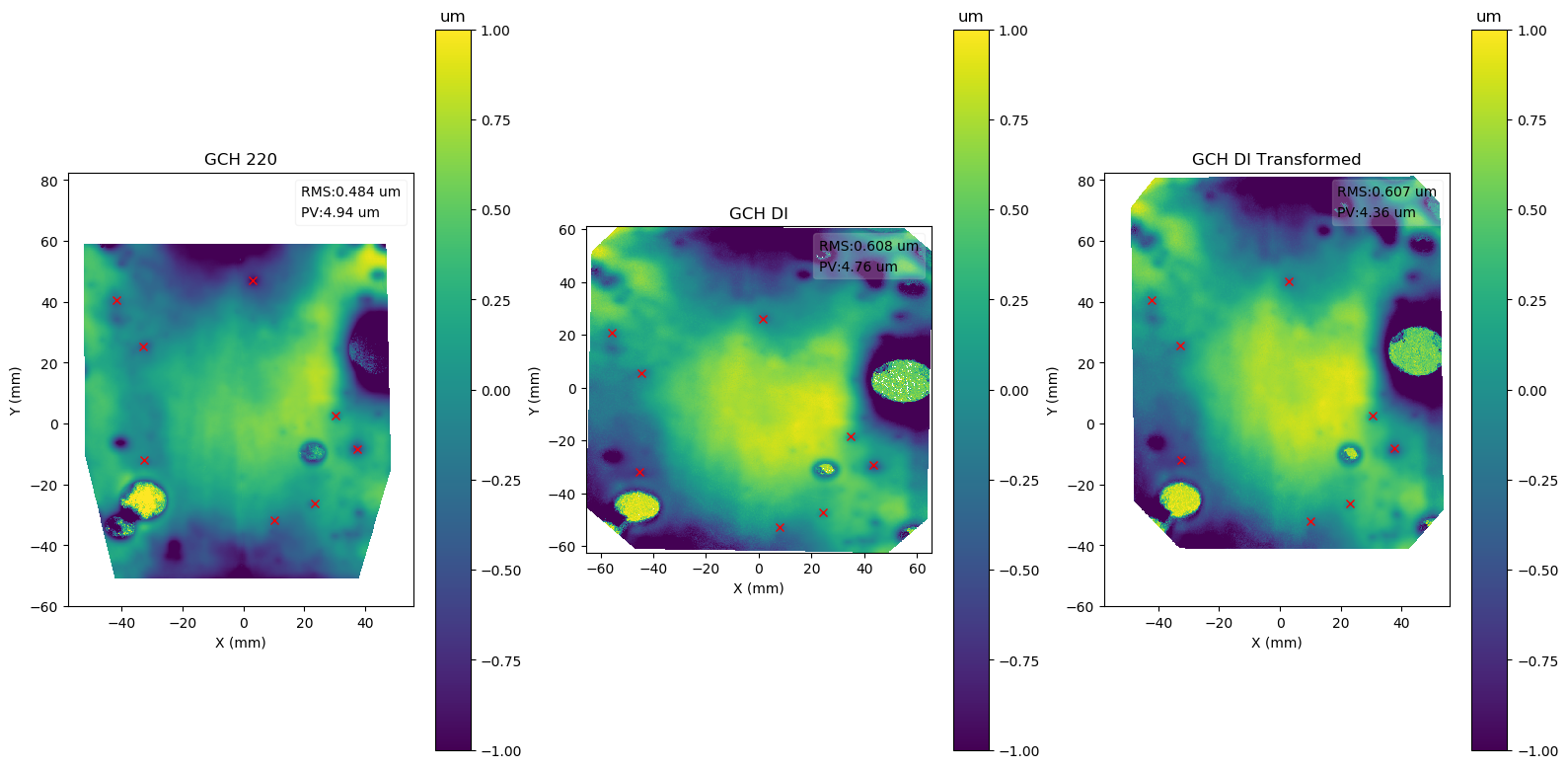
Methods .slope of object data2d_class.Data2D, and functions plot_slope_2D and plot_slope_slice calculate slopes and related statistics.
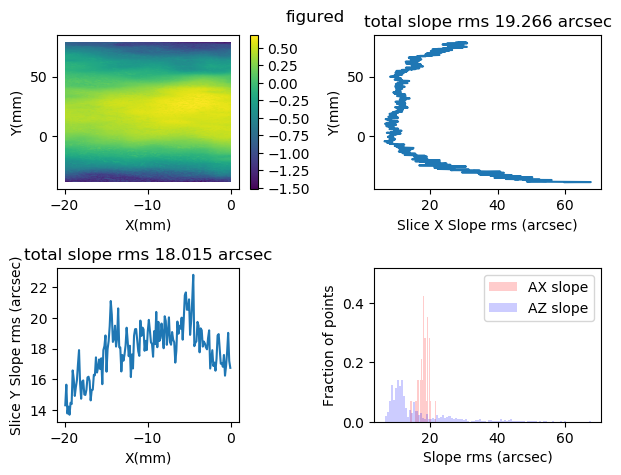
Data2D.slope?
Signature: Data2D.slope(self, *args, **kwargs) Docstring: slope_2D(wdata, x, y, scale=(1.0, 1.0, 1.0)) calculate slope maps in x and y. return a couple of maps of type slope,x,y data for x and y slopes respectively. Set scale to (dx,dy,1000.) for z in micron, x,y in mm. (?does it mean 1,1,1000?) File: c:\users\kovor\documents\python\pyxtel\source\pysurf\data2d_class.py Type: function
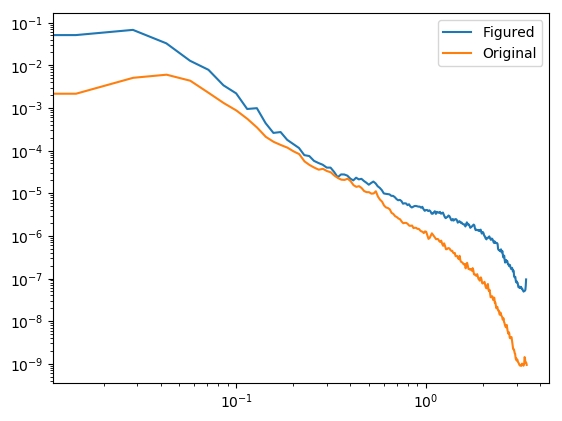
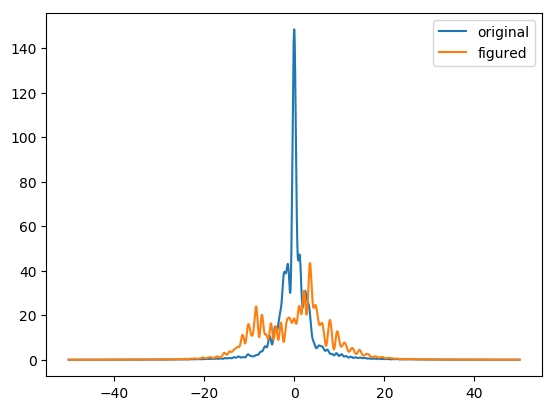
In this experiment, a part was measured before and after a process.
The two sets of data need to be aligned and subtracted, which can be done
 |  |
 |  |  |

Project page (docs):
Source Code: https://github.com/vincenzooo/pyXsurf
My email: vincenzo.cotroneo@inaf.it
Thanks to my current Institution (Osservatorio Astronomico di Brera, Merate, IT) and the previous one (Center for Astrophysics | Harvard & Smithsonian, Cambridge, MA, USA).
Thanks to the wonderful Python community.